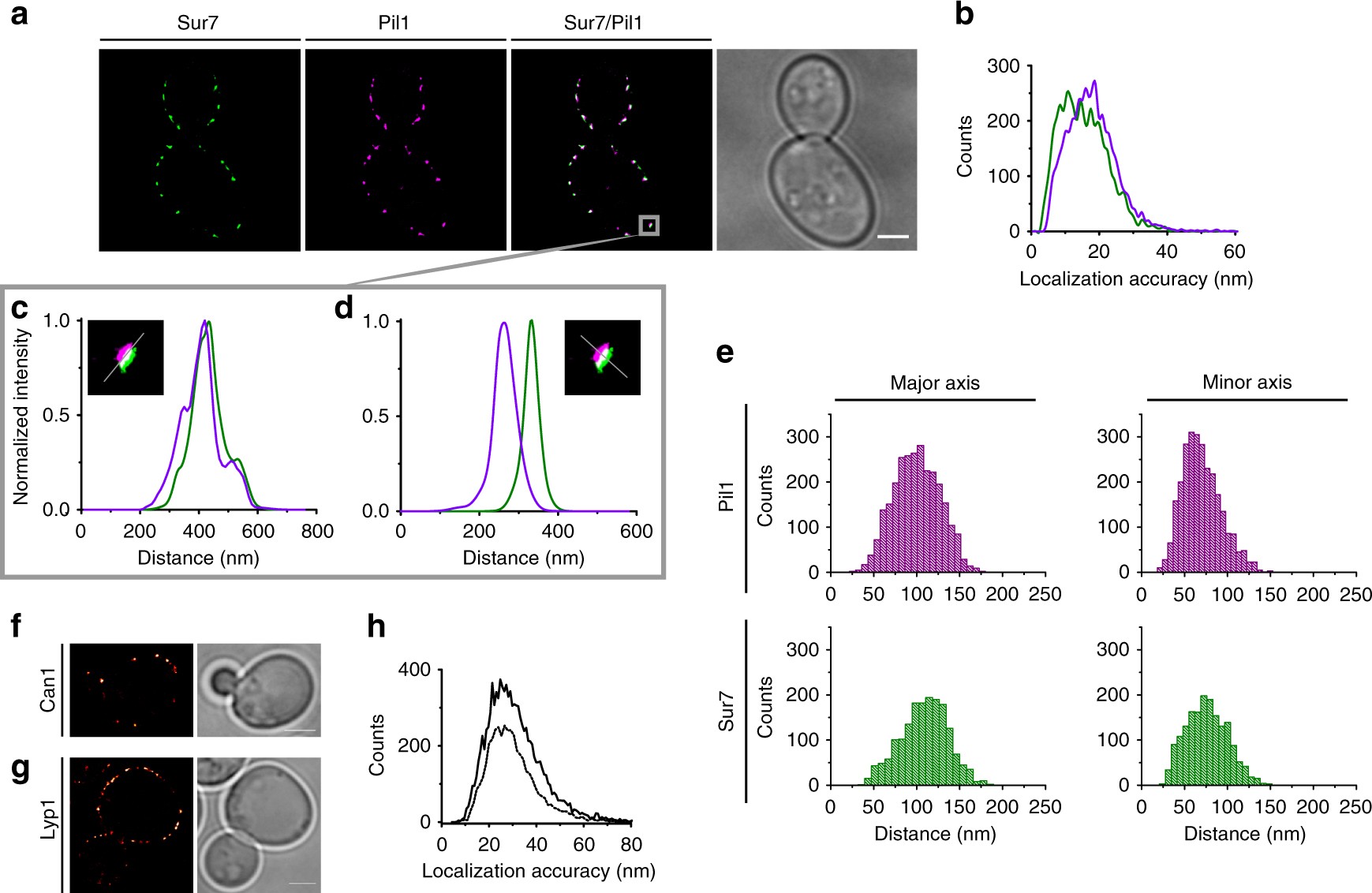
Steric exclusion and protein conformation determine the localization of plasma membrane transporters

Steric exclusion and protein conformation determine the localization of plasma membrane transporters

Steric exclusion and protein conformation determine the localization of plasma membrane transporters

Profiling Glycosylphosphatidylinositol (GPI)-Interacting Proteins in the Cell Membrane Using a Bifunctional GPI Analogue as the Probe
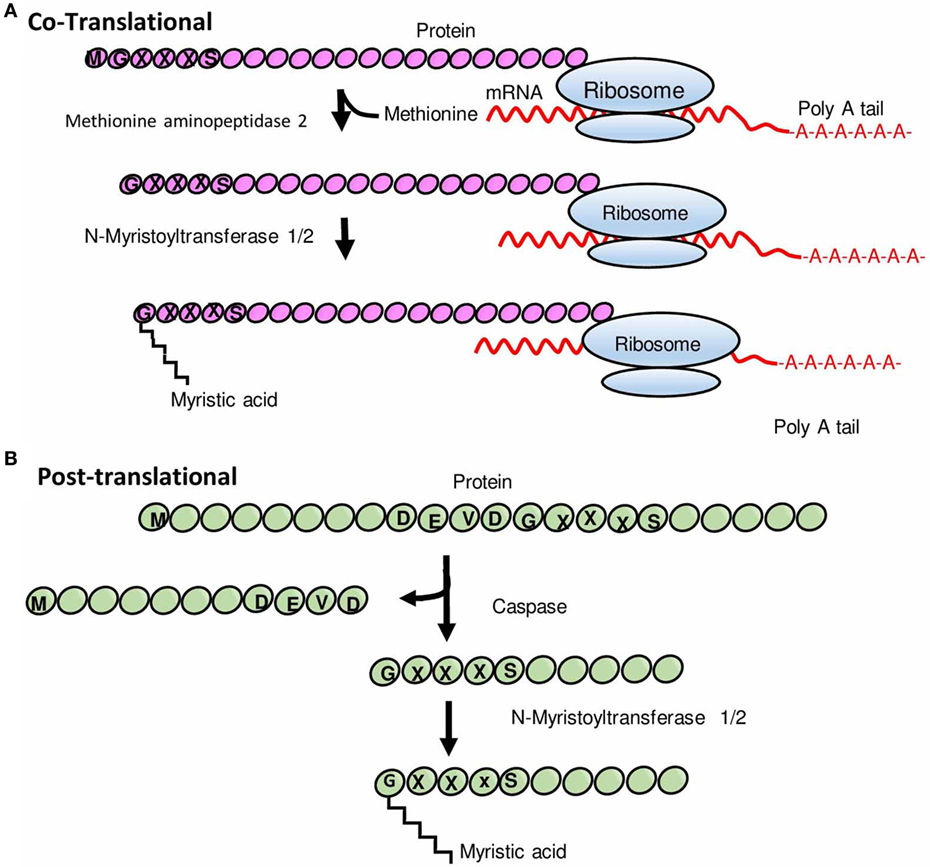
Frontiers Myristoylation: An Important Protein Modification in the Immune Response

Phospholipid distribution in the cytoplasmic membrane of Gram-negative bacteria is highly asymmetric, dynamic, and cell shape-dependent

Atlas of Subcellular RNA Localization Revealed by APEX-Seq - ScienceDirect
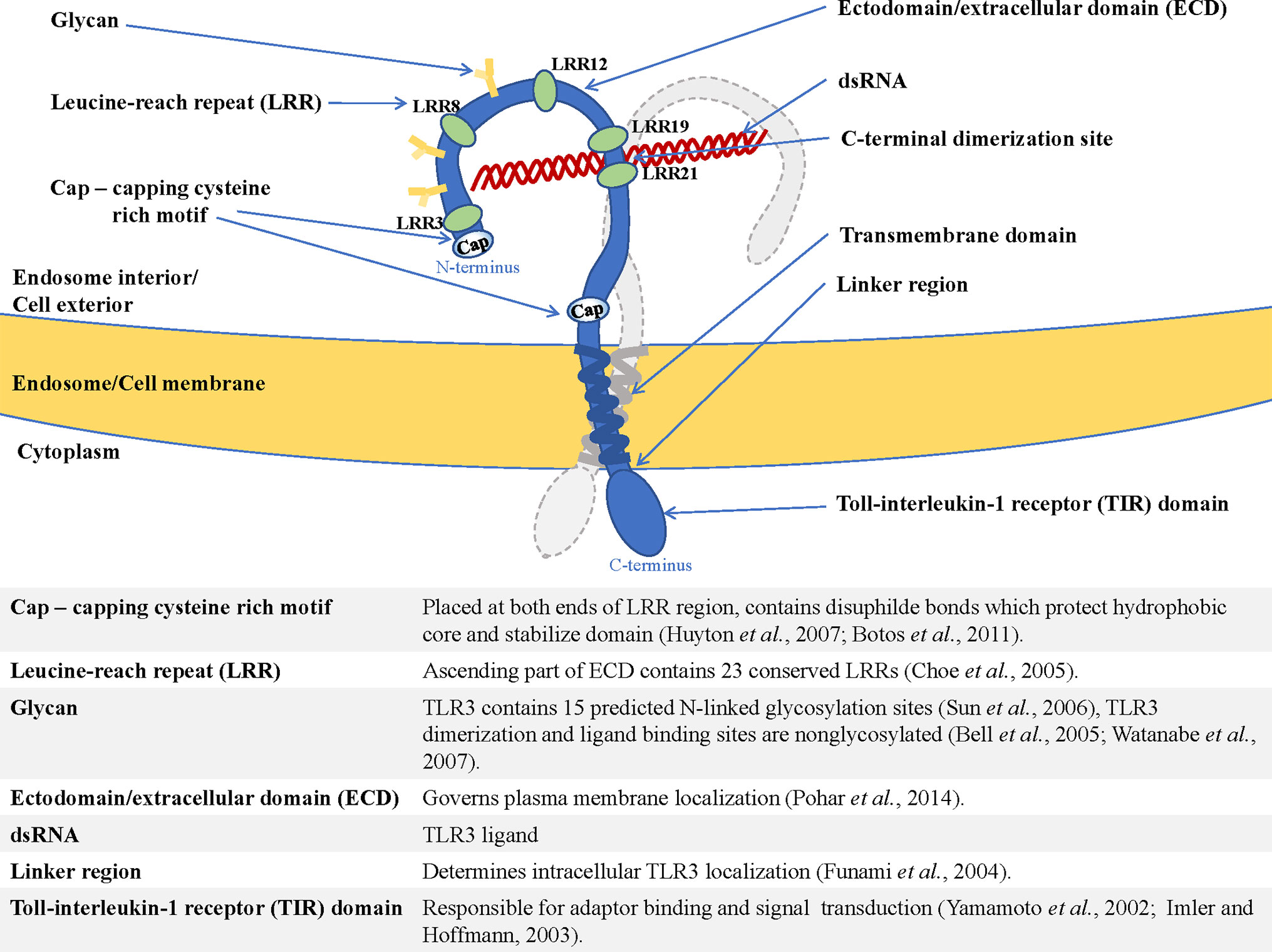
Frontiers Cell Surface Expression of Endosomal Toll-Like Receptors—A Necessity or a Superfluous Duplication?

Steric exclusion and protein conformation determine the localization of plasma membrane transporters

RNA localization in prokaryotes: Where, when, how, and why - Irastortza‐Olaziregi - 2021 - WIREs RNA - Wiley Online Library

Substrate-induced differential degradation and partitioning of the two tryptophan permeases Tat1 and Tat2 into eisosomes in Saccharomyces cerevisiae - ScienceDirect
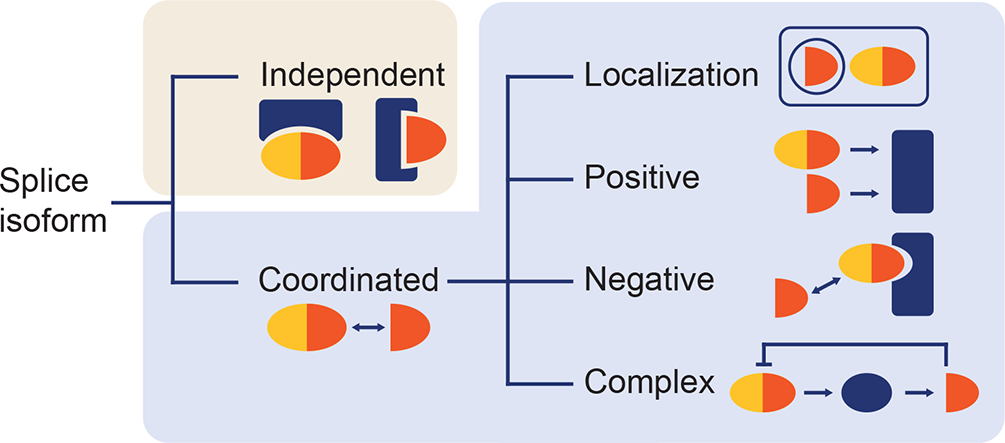
How alternative splicing changes the properties of plant proteins, Quantitative Plant Biology
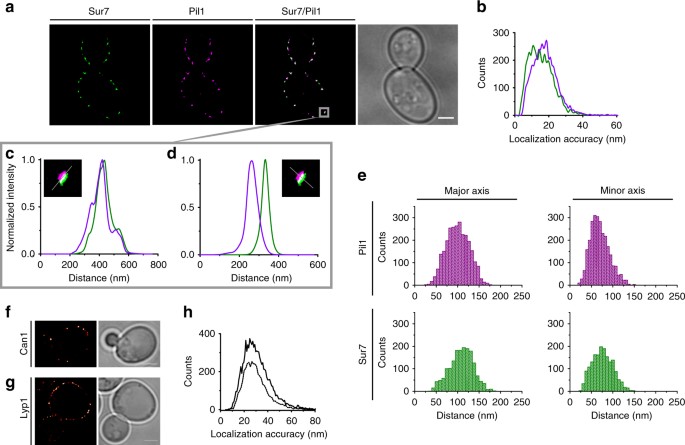
Steric exclusion and protein conformation determine the localization of plasma membrane transporters
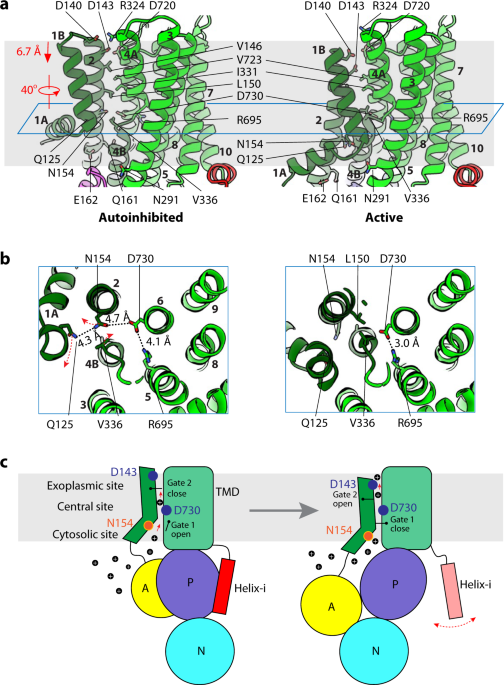
Structure and activation mechanism of the hexameric plasma membrane H+-ATPase

Spatial Proteomics Reveals Differences in the Cellular Architecture of Antibody-Producing CHO and Plasma Cell–Derived Cells - ScienceDirect